Human Kinome
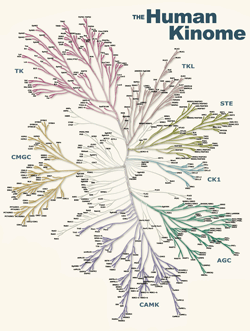
These pages support the publication of our report of the full complement of human protein kinases (the human kinome) in Science, on Dec 6, 2002:
The Protein Kinase Complement of the Human Genome
G Manning, DB Whyte, R Martinez, T Hunter, S Sudarsanam (2002).
Science 298:1912-1934
Science Abstract, Salk Institute Press Release, and free fulltext reprint provided by Science.
Additional material and links
A Human Kinome poster (1.9 Mb) accompanies the article, featuring a dendrogram of human protein kinases (see picture above). You can order a free printed copy from our colleagues at Cell Signaling Technology.
Science's STKE has commentary and related articles in a special kinome issue.
The classification, annotation and sequences of human, fly, worm and yeast kinases may now be explored through our KinBase database.
Kinase Phylogeny pages provide alignments and phylogenetic trees to show how human kinases relate to each other.
cDNA, protein and kinase domain sequences are available as FastA formatted text files, and may be searched using Blast.
Supplementary material for the paper is available as a set of compressed zip files at Science, or as individual files here:
- Tables in Excel format: Tables S1, S2, S3, S4, S5, S6, S7
- Tables in tab-delimited text format: Tables S1, S2, S3, S4, S5, S6, S7
- Table Legends (PDF)
- Materials and Methods (PDF)
- Notes on atypical protein kinase (aPK) families (PDF)
Update: (Nov 04): After the comparative analysis of mouse and human kinases, we found several kinases which were active in mouse but decayed to pseudogenes in human. For more information on these and other recently deceased human genes, see the page on gene death in the human lineage.
Update: (Dec 07): An updated table (Excel) includes Gene IDs, refseq accessions, HGNC names and improved sequences for some kinases.
Previous analysis of mammalian kinases
Here are some slightly dated diagrams showing the domain structure of kinases and phostphatases (PDF format)
- Mammalian receptor tyrosine kinases
- Mammalian cytoplasmic tyrosine kinases
- Mammalian protein tyrosine phosphatases
Here are other files made obsolete by our current analysis, retained for historical reasons:
- Unanchored dendrogram of mammalian protein kinases.
- Anchored dendrogram of mammalian protein kinases.
- Hyperbolic tree view of mammalian protein kinases
- Table of Aurora-family kinases (Excel97)
- Dendrogram of Aurora-family kinases
- Multiple alignment of Aurora-family kinases